okazi
Saturday, September 30, 2017
Wednesday, September 8, 2010
In translation
Messenger RNA (mRNA) carries information about a protein sequence to the ribosomes, the protein synthesis factories in the cell. It is coded so that every three nucleotides (a codon) correspond to one amino acid. In eukaryotic cells, once precursor mRNA (pre-mRNA) has been transcribed from DNA, it is processed to mature mRNA. This removes its introns—non-coding sections of the pre-mRNA. The mRNA is then exported from the nucleus to the cytoplasm, where it is bound to ribosomes and translated into its corresponding protein form with the help of tRNA. In prokaryotic cells, which do not have nucleus and cytoplasm compartments, mRNA can bind to ribosomes while it is being transcribed from DNA. After a certain amount of time the message degrades into its component nucleotides with the assistance of ribonucleases.[20]
Transfer RNA (tRNA) is a small RNA chain of about 80 nucleotides that transfers a specific amino acid to a growing polypeptide chain at the ribosomal site of protein synthesis during translation. It has sites for amino acid attachment and an anticodon region for codon recognition that binds to a specific sequence on the messenger RNA chain through hydrogen bonding.[25]
Ribosomal RNA (rRNA) is the catalytic component of the ribosomes. Eukaryotic ribosomes contain four different rRNA molecules: 18S, 5.8S, 28S and 5S rRNA. Three of the rRNA molecules are synthesized in the nucleolus, and one is synthesized elsewhere. In the cytoplasm, ribosomal RNA and protein combine to form a nucleoprotein called a ribosome. The ribosome binds mRNA and carries out protein synthesis. Several ribosomes may be attached to a single mRNA at any time.[20] rRNA is extremely abundant and makes up 80% of the 10 mg/ml RNA found in a typical eukaryotic cytoplasm.[27]
Transfer-messenger RNA (tmRNA) is found in many bacteria and plastids. It tags proteins encoded by mRNAs that lack stop codons for degradation and prevents the ribosome from stalling.[28]
[edit] Regulatory RNAs
In translation
Messenger RNA (mRNA) carries information about a protein sequence to the ribosomes, the protein synthesis factories in the cell. It is coded so that every three nucleotides (a codon) correspond to one amino acid. In eukaryotic cells, once precursor mRNA (pre-mRNA) has been transcribed from DNA, it is processed to mature mRNA. This removes its introns—non-coding sections of the pre-mRNA. The mRNA is then exported from the nucleus to the cytoplasm, where it is bound to ribosomes and translated into its corresponding protein form with the help of tRNA. In prokaryotic cells, which do not have nucleus and cytoplasm compartments, mRNA can bind to ribosomes while it is being transcribed from DNA. After a certain amount of time the message degrades into its component nucleotides with the assistance of ribonucleases.[20]
Transfer RNA (tRNA) is a small RNA chain of about 80 nucleotides that transfers a specific amino acid to a growing polypeptide chain at the ribosomal site of protein synthesis during translation. It has sites for amino acid attachment and an anticodon region for codon recognition that binds to a specific sequence on the messenger RNA chain through hydrogen bonding.[25]
Ribosomal RNA (rRNA) is the catalytic component of the ribosomes. Eukaryotic ribosomes contain four different rRNA molecules: 18S, 5.8S, 28S and 5S rRNA. Three of the rRNA molecules are synthesized in the nucleolus, and one is synthesized elsewhere. In the cytoplasm, ribosomal RNA and protein combine to form a nucleoprotein called a ribosome. The ribosome binds mRNA and carries out protein synthesis. Several ribosomes may be attached to a single mRNA at any time.[20] rRNA is extremely abundant and makes up 80% of the 10 mg/ml RNA found in a typical eukaryotic cytoplasm.[27]
Transfer-messenger RNA (tmRNA) is found in many bacteria and plastids. It tags proteins encoded by mRNAs that lack stop codons for degradation and prevents the ribosome from stalling.[28]
[edit] Regulatory RNAs
Ribonucleic acid (RNA) is a biologically important type of molecule that consists of a long chain of nucleotide units. Each nucleotide consists of a nitrogenous base, a ribose sugar, and a phosphate. RNA is very similar to DNA, but differs in a few important structural details: in the cell, RNA is usually single-stranded, while DNA is usually double-stranded; RNA nucleotides contain ribose while DNA contains deoxyribose (a type of ribose that lacks one oxygen atom); and RNA has the base uracil rather than thymine that is present in DNA. RNA is transcribed from DNA by enzymes called RNA polymerases and is generally further processed by other enzymes. RNA is central to protein synthesis. Here, a type of RNA called messenger RNA carries information from DNA to structures called ribosomes. These ribosomes are made from proteins and ribosomal RNAs, which come together to form a molecular machine that can read messenger RNAs and translate the information they carry into proteins. There are many RNAs with other roles – in particular regulating which genes are expressed, but also as the genomes of most viruses.
Contents[hide] |
[edit] Comparison with DNA
RNA and DNA are both nucleic acids, but differ in three main ways. First, unlike DNA which is double-stranded, RNA is a single-stranded molecule in most of its biological roles and has a much shorter chain of nucleotides. Second, while DNA contains deoxyribose, RNA contains ribose (there is no hydroxyl group attached to the pentose ring in the 2' position in DNA). These hydroxyl groups make RNA less stable than DNA because it is more prone to hydrolysis. Third, the complementary base to adenine is not thymine, as it is in DNA, but rather uracil, which is an unmethylated form of thymine.[1]
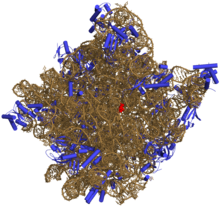
Like DNA, most biologically active RNAs, including mRNA, tRNA, rRNA, snRNAs and other non-coding RNAs, contain self-complementary sequences that allow parts of the RNA to fold and pair with itself to form double helices. Structural analysis of these RNAs has revealed that they are highly structured. Unlike DNA, their structures do not consist of long double helices but rather collections of short helices packed together into structures akin to proteins. In this fashion, RNAs can achieve chemical catalysis, like enzymes.[2] For instance, determination of the structure of the ribosome—an enzyme that catalyzes peptide bond formation—revealed that its active site is composed entirely of RNA.[3]
[edit] Structure
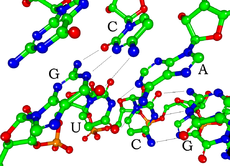
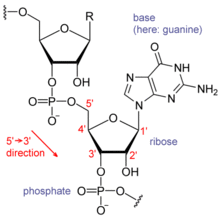
Each nucleotide in RNA contains a ribose sugar, with carbons numbered 1' through 5'. A base is attached to the 1' position, generally adenine (A), cytosine (C), guanine (G) or uracil (U). Adenine and guanine are purines, cytosine and uracil are pyrimidines. A phosphate group is attached to the 3' position of one ribose and the 5' position of the next. The phosphate groups have a negative charge each at physiological pH, making RNA a charged molecule (polyanion). The bases may form hydrogen bonds between cytosine and guanine, between adenine and uracil and between guanine and uracil.[4] However other interactions are possible, such as a group of adenine bases binding to each other in a bulge,[5] or the GNRA tetraloop that has a guanine–adenine base-pair.[4]
An important structural feature of RNA that distinguishes it from DNA is the presence of a hydroxyl group at the 2' position of the ribose sugar. The presence of this functional group causes the helix to adopt the A-form geometry rather than the B-form most commonly observed in DNA.[6] This results in a very deep and narrow major groove and a shallow and wide minor groove.[7] A second consequence of the presence of the 2'-hydroxyl group is that in conformationally flexible regions of an RNA molecule (that is, not involved in formation of a double helix), it can chemically attack the adjacent phosphodiester bond to cleave the backbone.[8]
RNA is transcribed with only four bases (adenine, cytosine, guanine and uracil),[9] but there are numerous modified bases and sugars in mature RNAs. Pseudouridine (Ψ), in which the linkage between uracil and ribose is changed from a C–N bond to a C–C bond, and ribothymidine (T), are found in various places (most notably in the TΨC loop of tRNA).[10] Another notable modified base is hypoxanthine, a deaminated adenine base whose nucleoside is called inosine (I). Inosine plays a key role in the wobble hypothesis of the genetic code.[11] There are nearly 100 other naturally occurring modified nucleosides,[12] of which pseudouridine and nucleosides with 2'-O-methylribose are the most common.[13] The specific roles of many of these modifications in RNA are not fully understood. However, it is notable that in ribosomal RNA, many of the post-transcriptional modifications occur in highly functional regions, such as the peptidyl transferase center and the subunit interface, implying that they are important for normal function.[14]
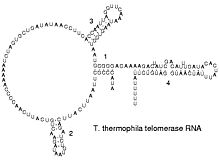
The functional form of single stranded RNA molecules, just like proteins, frequently requires a specific tertiary structure. The scaffold for this structure is provided by secondary structural elements which are hydrogen bonds within the molecule. This leads to several recognizable "domains" of secondary structure like hairpin loops, bulges and internal loops.[15] Since RNA is charged, metal ions such as Mg2+ are needed to stabilise many secondary and tertiary structures.[16]
[edit] Synthesis
Synthesis of RNA is usually catalyzed by an enzyme—RNA polymerase—using DNA as a template, a process known as transcription. Initiation of transcription begins with the binding of the enzyme to a promoter sequence in the DNA (usually found "upstream" of a gene). The DNA double helix is unwound by the helicase activity of the enzyme. The enzyme then progresses along the template strand in the 3’ to 5’ direction, synthesizing a complementary RNA molecule with elongation occurring in the 5’ to 3’ direction. The DNA sequence also dictates where termination of RNA synthesis will occur.[17]
RNAs are often modified by enzymes after transcription. For example, a poly(A) tail and a 5' cap are added to eukaryotic pre-mRNA and introns are removed by the spliceosome.
There are also a number of RNA-dependent RNA polymerases that use RNA as their template for synthesis of a new strand of RNA. For instance, a number of RNA viruses (such as poliovirus) use this type of enzyme to replicate their genetic material.[18] Also, RNA-dependent RNA polymerase is part of the RNA interference pathway in many organisms.[19]
[edit] Types of RNA
[edit] Overview

Messenger RNA (mRNA) is the RNA that carries information from DNA to the ribosome, the sites of protein synthesis (translation) in the cell. The coding sequence of the mRNA determines the amino acid sequence in the protein that is produced.[20] Many RNAs do not code for protein however (about 97% of the transcriptional output is non-protein-coding in eukaryotes [21][22][23][24]).
These so-called non-coding RNAs ("ncRNA") can be encoded by their own genes (RNA genes), but can also derive from mRNA introns.[25] The most prominent examples of non-coding RNAs are transfer RNA (tRNA) and ribosomal RNA (rRNA), both of which are involved in the process of translation.[1] There are also non-coding RNAs involved in gene regulation, RNA processing and other roles. Certain RNAs are able to catalyse chemical reactions such as cutting and ligating other RNA molecules,[26] and the catalysis of peptide bond formation in the ribosome;[3] these are known as ribozymes.
[edit] In translation
Messenger RNA (mRNA) carries information about a protein sequence to the ribosomes, the protein synthesis factories in the cell. It is coded so that every three nucleotides (a codon) correspond to one amino acid. In eukaryotic cells, once precursor mRNA (pre-mRNA) has been transcribed from DNA, it is processed to mature mRNA. This removes its introns—non-coding sections of the pre-mRNA. The mRNA is then exported from the nucleus to the cytoplasm, where it is bound to ribosomes and translated into its corresponding protein form with the help of tRNA. In prokaryotic cells, which do not have nucleus and cytoplasm compartments, mRNA can bind to ribosomes while it is being transcribed from DNA. After a certain amount of time the message degrades into its component nucleotides with the assistance of ribonucleases.[20]
Transfer RNA (tRNA) is a small RNA chain of about 80 nucleotides that transfers a specific amino acid to a growing polypeptide chain at the ribosomal site of protein synthesis during translation. It has sites for amino acid attachment and an anticodon region for codon recognition that binds to a specific sequence on the messenger RNA chain through hydrogen bonding.[25]
Ribosomal RNA (rRNA) is the catalytic component of the ribosomes. Eukaryotic ribosomes contain four different rRNA molecules: 18S, 5.8S, 28S and 5S rRNA. Three of the rRNA molecules are synthesized in the nucleolus, and one is synthesized elsewhere. In the cytoplasm, ribosomal RNA and protein combine to form a nucleoprotein called a ribosome. The ribosome binds mRNA and carries out protein synthesis. Several ribosomes may be attached to a single mRNA at any time.[20] rRNA is extremely abundant and makes up 80% of the 10 mg/ml RNA found in a typical eukaryotic cytoplasm.[27]
Transfer-messenger RNA (tmRNA) is found in many bacteria and plastids. It tags proteins encoded by mRNAs that lack stop codons for degradation and prevents the ribosome from stalling.[28]